
Welcome to the Expitope server!
It enables you to search for potential cross-reactions of your T cell therapy targets.
After submitting an epitope and an allowed number of mismatches you will obtain
all proteins which contain the given peptide or variants thereof. The expression values of the corresponding transcripts in various tissues will be returned.
Additionally, Expitope will provide you with the probability that your peptide will be created by proteasomal cleavage, that it will bind to the TAP transporter and its affinity to MHC class I alleles.
Submit Expitope job
Usage
Expitope server only requires your T cell therapy target epitope and the maximum number of mismatches up to which you want to consider potential off-target effects.
We provide all known human proteins containing the exact or approximate sequence, together with their associated transcripts. Additionally,
Expitope returns normalised expression values for all these identified transcripts in 16 healthy tissues and three positive control cancer cell lines. To aid the evalution of the found off-target hits with regard to potential cross-reactivity, we score all results based on affinity to immunesystem components.
The basis for Expitope's analysis builds a RNA-Seq database compiled from ENCODE (GEO identifier: GSM758575, GSM981253 and GSM958749) and Illumina Human Body Map data (GEO identifier: GSE30611). The latter provides transcriptome data for 16 normal human tissues, from the former we chose three cancerous cell lines as positive control.
We extracted the read counts from the sequencing data for all GenCodeV19 transcripts and normalised them to FPKM values (Fragments Per Kilobase of exon per Million fragments mapped).
Additionally, we map the found proteins on the results of a transcriptome analysis by Wang et al. (2008), who provide RPKM values for 23,115 Ensembl gene identifier of 15 different human tissues, among them six independent brain samples, which are of particular importance for avoiding cross-reactivity.
The identified hits are scored according to their probability of being created by proteasomal cleavage and their affinities to the TAP transporter and MHC complex. For this purposes, we applied NetChop 3.1, a matrix approach from Peters et al. (2003) and the portable version of NetMHC 3.0, respectively. From this three components a combined score is calculated by which the list of results is sorted and presented to the user.

Input
Epitope and Mismatches
Please note that the amino acid sequence has to have a length of at least seven positions in order to avoid excessive database matches. The number of mismatches cannot exceed a half of the size of the provided epitope for the same reason. Furthermore, MHC binding scores can only be provided for epitopes with a length between 8 and 14 positions, due to the implementation of the underlying software.
When you want to define fixed positions in the epitope input sequence, check the corresponding box and provide the sequence in a case sensitive manner. E.g., KvaeLvhfL defines a nonamer where the first, fifth and last position can not be modified.
Proteasomal Cleavage Prediction
When calculating the cleavage probability for the current epitope, we followed the original paper by Keşmir et al. (2002) and used the formula

wherein Pc is the probability that the peptide is cleaved exactly at the C-terminus and Pcon represents the probability of the rest of the peptide staying intact:
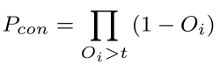
Due to the overall cleavage probability being a product, it becomes very small for longer input sequences. Hence it is advisable to only rely on this score for peptides in the range of seven to eleven amino acids, as that is the epitope size for which NetChop has been most extensively tested, although the calculation itself does not limit the input to a certain length.
TAP Affinity Prediction
When testing the divergence between predicted and experimentally tested IC50 values, the authors concluded that the best concordance is achieved when taking precursor peptides into account, i.e. instead of the initial nonamer they calculated the affinity for an N-terminal elongated sequence. As the length of the epitopes submitted to our server is defined by the users, we modified the established formula to work without precursor sequences:

where Ni denotes the i-th amino acid from the N-terminus of the given peptide and α is a weight that determines the influence of the N-terminal residues on the overall affinity score. Hence, only the IC50 values for the C-terminal residue as well as a weighted sum of the three N-terminal amino acids are used for the scoring. The authors experimentally determined the best values for weight α to be 0.2, but it can be changed in the Expitope web server by the user into another floating point number.
Although it is technically possible to score peptides of any length ≥ 4 with this approach, it has to be kept in mind that the matrices are constructed on the basis of nonamer epitopes and have also only been extensively tested one those or with potential precursors. When analyzing longer peptides the returned values might not reflect the real affinity to TAP and it could be beneficial to exclude the N terminus in such cases.
MHC Binding Affinity
Combined Score

where P is the proteasomal cleavage probability and the A-terms are affinities in IC50 values to the transporter associated with antigen processing (TAP) and the MHC complex as explained above.
Output

The output is provided as a tab-separated text file for download and as a formatted table for data inspection by eye. It is sorted by ascending number of mismatches and descending combined score.
The columns denote the RefSeq protein identifier to which the matching peptide belongs and the exact sequence that matched the input parameters. The column titled index gives the starting position of the epitope in the full length RefSeq entry. The following columns denote number of mismatches which separate the current match from the input sequence and the three scores corresponding to cleavage probability, TAP affinity and MHC affinity as well as the resulting combined score.
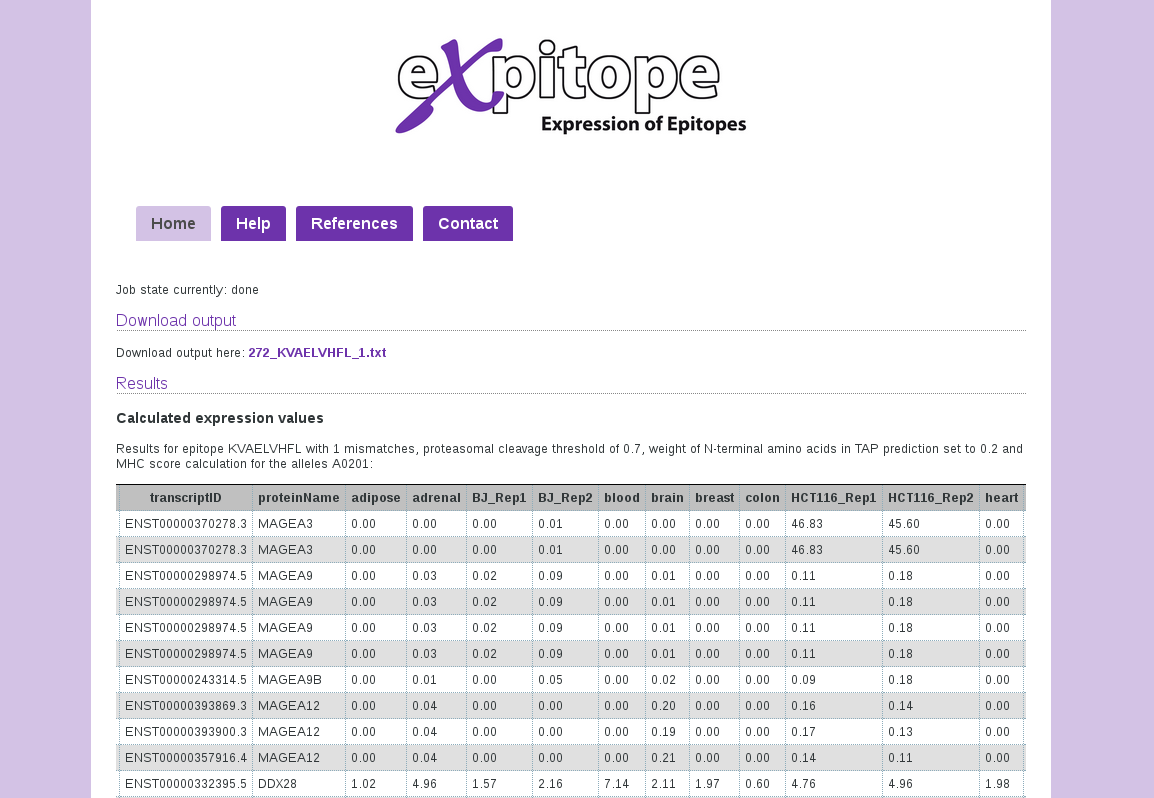
The second part of the results table contains information regarding the transcripts and their expression. Expitope first lists the ENSEMBL transcript identifier corresponding to the protein, its official name, followed by the RPKM values in all analysed tissues and cell lines (alphabetical order). The results may contain the same transcript ID for different protein IDs or a protein ID can be associated with multiple transcript IDs.
The results based on data from Wang et al. (2008) are presented in an extra table, following the same format as described above. Additionally, Expitope lists proteins, which contain the provided epitope but could not be matched to a transcript identifier. These are usually automatically determined proteins (recognizable through their "XP_" identifier start instead of "NP_") whose real existence has not yet been confirmed.
References
If you'd like to learn more about the data and scoring system used by Expitope, please consider the following original publications:
NetChop
The role of the proteasome in generating cytotoxic T cell epitopes: Insights obtained from improved predictions of proteasomal cleavage.M. Nielsen, C. Lundegaard, O. Lund, and C. Keşmir. Immunogenetics, 57(1-2):33-41, 2005. PMID:15744535 Prediction of proteasome cleavage motifs by neural networks.
C. Keşmir, A. Nussbaum, H. Schild, V. Detours, and S. Brunak. Prot. Eng., 15(4): 287-296, 2002. PMID:11983929
TAP affinity
Identifying MHC Class I Epitopes by Predicting the TAP Transport Efficiency of Epitope Precursors. B. Peters, S. Bulik, R. Tampe, P. M. van Endert, H-G. Holzhütter. J Immunol, 171:1741-1749, 2003. PMID:12902473NetMHC
Reliable prediction of T-cell epitopes using neural networks with novel sequence representations. Nielsen M, Lundegaard C, Worning P, Lauemøller SL, Lamberth K, Buus S, Brunak S, Lund O. Protein Sci., 12:1007-17, 2003. PMID:12717023 NetMHC-3.0: accurate web accessible predictions of human, mouse and monkey MHC class I affinities for peptides of length 8-11 Lundegaard C, Lamberth K, Harndahl M, Buus S, Lund O, Nielsen M. Nucleic Acids Res. 1;36(Web Server issue):W509-12. 2008. PMID:18463140 Accurate approximation method for prediction of class I MHC affinities for peptides of length 8, 10 and 11 using prediction tools trained on 9mers. Lundegaard C, Lund O, Nielsen M. Bioinformatics, 24(11):1397-98, 2008. PMID:18413329Expression data
An integrated encyclopedia of DNA elements in the human genome. The ENCODE Project Consortium. Nature, 489(7414):57-74, 2012. PMID:22955616Alternative isoform regulation in human tissue transcriptomes.
Wang ET, Sandberg R, Luo S, Khrebtukova I, Zhang L, Mayr C, Kingsmore SF, Schroth GP, Burge CB. Nature 456:470-476, 2008. PMID:18978772
Maintenance
This web site is updated and maintained by Kerstin Haase. If you encounter any problems or have comments regarding Expitope, please contact haase[at]wzw.tum.de.
Collaboration

The Expitope web server was developed in collaboration with Medigene Immunotherapies GmbH a subsidiary of Medigene AG.
Working Group
This service is hosted by the Wissenschaftszentrum Weihenstephan, Department of Genome-Oriented Bioinformatics of the Technical University Munich.
Expitope was designed and created by members of the Frishman group.
Disclaimer
No warranties or guaranties concerning this web based tool "Expitope", express or implied, including but not limited to a warranty of merchantability or fitness for a particular purpose is given. This web based tool "Expitope" is solely intended to be an initial source of information in TCR selection based on certain assumptions that may or may not align with your specific conditions, without the warranty or guaranty of generated data to be complete, reliable and externally verified. This web based tool "Expitope" does not provide any warranty or guaranty regarding toxicology data and information in animals or humans and does not provide any data and information regarding protein sequences.
No warranties or guaranties are given that the generated data and information will meet your requirements or operate under your specific conditions of use. In addition, no warranties or guaranties are given that use of this web based tool "Expitope" will be secure, error free, or free from interruption. The user shall determine upon its sole responsibility whether generated data and information sufficiently meets any needs and requirements. The user is solely responsible and liable for any loss incurred due to failure of this web based tool "Expitope" to meet the user’s requirements. No liability is given to the user or any other third party for indirect, consequential, special or punitive damages resulting from the use of the web based tool "Expitope".